Diagnostic Evaluation of Non-Interpretable Results Associated with rpoB Gene in Genotype MTBDRplus Ver 2.0
Article information
Abstract
Background
Line probe assay (LPA) is standard diagnostic tool to detect multidrug resistant tuberculosis. Non-interpretable (NI) results in LPA (complete missing or light wild-type 3 and 8 bands with no mutation band in rpoB gene region) poses a diagnostic challenge.
Methods
Sputum samples obtained between October 2016 and July 2017 at the Intermediate Reference Laboratory, All India Institute of Medical Sciences Hospital, New Delhi, India were screened. Smear-positive and smear-negative culture-positive specimens were subjected to LPA Genotype MTBDRplus Ver 2.0. Smear-negative with culture-negative and culture contamination were excluded. LPA NI samples were subjected to phenotypic drug susceptibility testing (pDST) using MGIT-960 and sequencing.
Results
A total of 1,614 sputum specimens were screened and 1,340 were included for the study (smear-positive [n=1,188] and smear-negative culture-positive [n=152]). LPA demonstrated 1,306 (97.5%) valid results with TUB (Mycobacterium tuberculosis) band, 24 (1.8%) NI, three (0.2%) valid results without TUB band, and seven (0.5%) invalid results. Among the NI results, 22 isolates (91.7%) were found to be rifampicin (RIF) resistant and two (8.3%) were RIF sensitive in the pDST. Sequencing revealed that rpoB mutations were noted in all 22 cases with RIF resistance, whereas the remaining two cases had wild-type strains. Of the 22 cases with rpoB mutations, the most frequent mutation was S531W (n=10, 45.5%), followed by S531F (n=6, 27.2%), L530P (n=2, 9.1%), A532V (n=2, 9.1%), and L533P (n=2, 9.1%).
Conclusion
The present study showed that the results of the Genotype MTBDRplus assay were NI in a small proportion of isolates. pDST and rpoB sequencing were useful in elucidating the cause and clinical meaning of the NI results.
Introduction
The high morbidity and mortality associated with drug resistant tuberculosis (DR-TB) makes it a matter of grave concern [1]. The widespread availability of rapid molecular diagnostic techniques has improved the case detection rate for DR-TB [2]. World Health Organization (WHO) approved the first version of line probe assay (Genotype MTBDRplus Ver 1.0, Hain Lifescience, Nehren, Germany [LPA V.1]) in 2008. The utility was limited to higher grade of smear-positive and culture-positive samples [3]. An improved version of line probe assay (LPA) (Genotype MTBDRplus Ver 2.0, Hain Lifescience [LPA V.2]) was endorsed by WHO in 2011 [4] which has high sensitivity and specificity when compared to Xpert MTB/RIF [5,6].
DNA based molecular diagnostics are based on screening alterations in the nucleic acid sequences. The principle of LPA is based on rpoB gene, an 81 base-pair rifampicin (RIF) resistance-determining region which accounts about 95% of RIF resistance. Missing of wild-type (WT) bands with or without presence of mutation (MUT) bands is interpreted as resistant. RIF specific mutation, S531L, H26Y, H526D, and D516V are the most prevalent mutation found in rpoB gene in LPA [2,7]. Disputed mutations have also been observed at positions L511P, D516Y, H526L, and L533P in rpoB gene [8,9]. In LPA V.2, a strain having missing or light band of WT3 (codon 513 to 517) or WT8 (codon 530 to 533) without any MUT band, could be due to silent mutation [7] and failure of hybridization probe [10]. Non-interpretable (NI) results in LPA V.2., albeit a small proportion, are a diagnostic challenge. Our study evaluated the role of phenotypic drug susceptibility testing (pDST) and sequencing in NI results of LPA V.2.
Materials and Methods
1. Subject recruitment
Sputum samples were received at the Intermediate Reference Laboratory (IRL) between October 2016 to July 2017 from the directly observed treatment short-course centers and medical out-patient department of All India Institute of Medical Sciences Hospital (AIIMS), New Delhi, India were evaluated for inclusion in the study. Smear-positive and smear-negative culture-positive specimens were included in the study. Smear-negative with culture-negative and culture contamination were excluded.
Investigations were conducted at the IRL of the Department of Medicine at the AIIMS, New Delhi, India. The sputum specimens were processed in bio-safety cabinet II in a bio-safety level-3 laboratory. As per laboratory protocol, sputum samples of all tuberculosis (TB) suspects were subjected to smear microscopy by Ziehl-Neelsen (ZN) staining and Mycobacterial Growth Indicator Tube (MGIT)-960 liquid culture (LC). Before decontamination, sputum samples were subjected to direct smear microscopy examination by the ZN method. The samples were decontaminated using the NALC-NaOH method (final NaOH concentration, 1%). The decontaminated samples were then subjected to LPA and inoculated in LC and indirect drug susceptibility testing was performed by using the final critical concentration of anti-TB drugs, i.e., 0.1 μg/mL of isoniazid (INH) and 1.0 μg/mL of RIF. Immunochromatographic test was done on all culture-positive samples using immunochromatographic assay kit (SD MPT64TB Ag kit, Standard Diagnostics, Suwon, Korea).
LPA was done using LPA kit (Genotype MTBDRplus Ver 2.0, Hain Lifescience) and the results were interpreted based on the operating manual provided by the manufacturer, valid results with TUB (Mycobacterium tuberculosis) band: presence of conjugate control (CC) and amplification control (AC) bands with all loci bands of WT and/or MUT bands; valid results without TUB band: presence of only CC and AC bands (without WT and MUT bands); NI: presence of TUB band and all loci bands with completely missing or light band of WT3 or WT8 without any MUT band in rpoB gene; invalid: presence of TUB band without control probes [7]. LPA NI samples were subjected to pDST using MGIT-960 and sequencing.
Sanger based sequencing was used as a reference standard for this study. Sequencing was performed for rpoB gene (forward primer 5’-CAGACGTTGATCAACATCCG-3’ and reverse primer 5’-TACGGCGTTTCGATGAAC-3’) [11] on ABI prism 3130xl genetic analyzer (Applied Biosystems and Hitachi, Ltd., Carlsbad, CA, USA).
2. Statistical analysis
The nucleotide sequence chromatograms were analyzed using BioEdit Software (Thomas Hall, Raleigh, NC, USA) and ClustalW 2.0. Data was entered and analyzed in Microsoft Excel. No patient particulars were recorded.
3. Ethical approval
The study was conducted in samples collected under the national program with no patient interviews. Ethics approval was thus not required.
Results
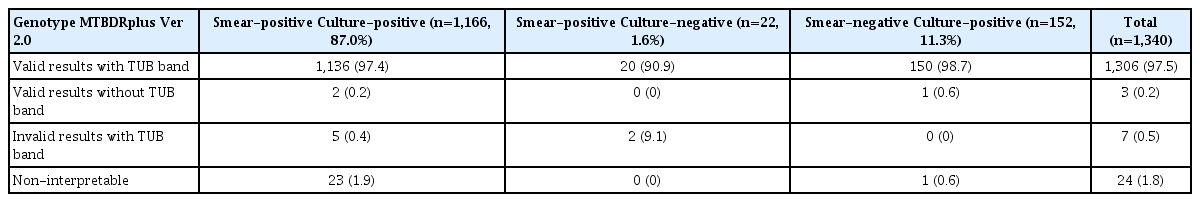
Out of total 1,614 sputum specimens, 1,212 (75.1%) were smear-positive and 402 (24.9%) were found to be smear-negative using ZN staining and microscopy. Among smear-positive specimens, 1,166 of 1,212 (96.2%) were culture-positive, 22 of 1,212 (1.8%) were culture-negative and contamination was found in 24 of 1,212 (2%) specimens. Among smear-negative specimens, 152 of 402 (37.8%) were culture-positive, 228 of 402 (56.7%) were culture-negative and rest 22 of 402 (5.5%) were found to be contaminated in LC (Table 1). After satisfying the inclusion and exclusion criteria, 1,340 samples were included for the study. All smear-positive specimens were directly processed for the LPA while, in case of smear-negative specimens, LPA were done only when the culture turned positive in LC (indirect).
1. Performance of line probe assay (LPA V.2)
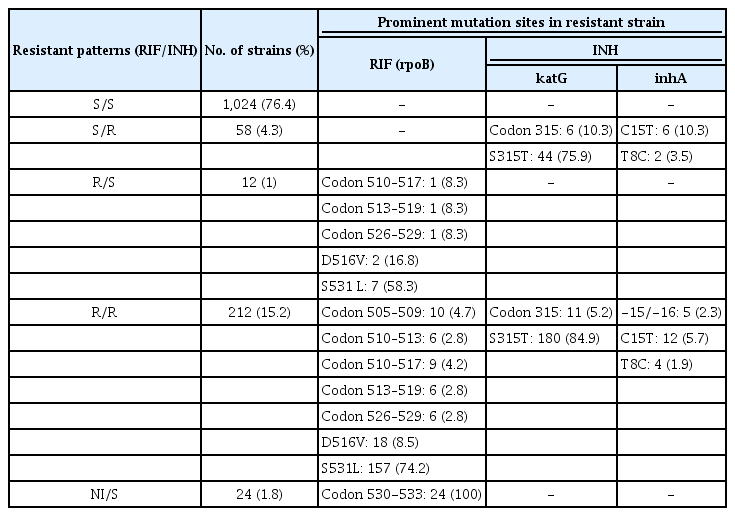
LPA detected 1,306 of 1,340 valid results with TUB band (97.5%) and 24 of 1,340 NI results (1.8%) (Table 1). Among valid results with TUB band, 1,024 of 1,340 specimens (76.4%) were observed to be sensitive to both RIF and INH drugs, 58 of 1,340 specimens (4.4%) were INH mono-resistant, 12 of 1,340 (1%) were RIF mono-resistant, multidrug resistant (MDR)-TB were found in 212 of 1,340 (15.8%) (Table 2).
The most frequent mutation in RIF-resistant isolates was S531L substitution (n=7, 58.3% in RIF mono-resistant strain and n=157, 74.2% in MDR-TB strain) in rpoB gene, followed by at position D516V (n=2, 16.8% in RIF mono-resistant strain and n=18, 8.5% in MDR-TB strain). Among INH resistant strain, katG gene was found to be the most prominent mutated gene at site S315T (n=44, 75.9% strains belong to INH mono-resistant and n=180, 84.9% were observed from MDR-TB strains), whereas in inhA gene, C15T mutation site was the most common in both INH mono-resistant (n=6, 10.3%) and MDR-TB strains (n=12, 5.7%). NI strains were observed in 24 specimens (1.8%) and were further evaluated with pDST and DNA sequencing.
2. Comparison of mutation patterns in NI with LPA, pDST, and sequencing
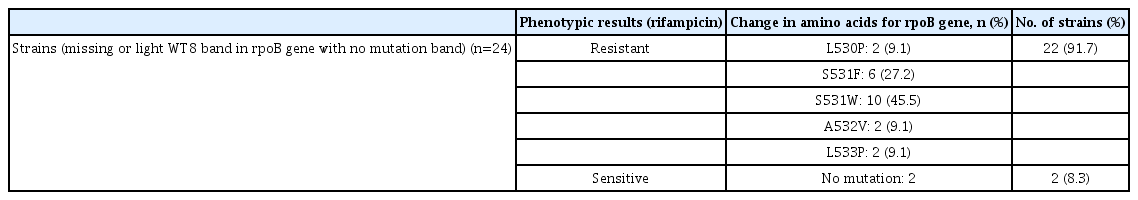
For the evaluation of mutation patterns, all 24 NI strains (1.8%) were put on pDST with standard critical concentration on both RIF and INH drugs. Among all NI, 22 (91.7%) were resistant and two (8.3%) were sensitive to RIF in pDST. Genetic analysis of rpoB gene revealed five unique mutations that were not mentioned by the manufacturer in the operating manual for the Genotype MTBDRplus assay. Among all unique mutations, S531W substitution was found in 10 strains (45.5%) followed by S531F substitution in six strains (27.2%), L530P, A532V, and L533P substitutions equally distributed in two strains each (Table 3).
Discussion
In last decade, the use of several molecular diagnostic techniques like Xpert MTB/RIF and LPA have increased the DR-TB case notification rate worldwide [12]. RIF-resistant (RR) associated rpoB gene hotspot (codon 507–533) is identified as the principle RR-TB defining region in all the nucleic acid amplification tests based molecular diagnostics [13].
We found that a low proportion of LPA V.2 results are NI (n=24, 1.8%). Silent mutations (WT3 [codon 514] and WT8 [codon 533]) and rare rpoB gene mutations [7,12] have been previously identified to cause NI results. In the recent version of LPA, mutation in WT8 band (codon 530–533) is a common cause of the NI result. In present study, rpoB associated WT8 band was completely missing without presence of any MUT bands in the strips (defined as NI) thereby leading to diagnostic dilemma. As per the programmatic guidelines in India, all RIF mono-resistant patients are eligible for treatment as MDR patient [1]. This limitation associated with LPA V.2 may lead to patients either misdiagnosed as RIF-sensitive TB resulting in high failure and relapse rate or RR-TB resulting in unnecessarily being treated as MDR-TB [14,15].
In this study, the most frequent mutation in rpoB coded gene was found at codon S531L in both RIF mono-resistant (n=7, 58.3%) as well as in MDR-TB isolates (n=157, 74.2%) which is consistent with the previously published studies [6,16,17]. For the detection of INH resistant, LPA is equipped with two genes katG and inhA, both of them showed similar frequency of mutation as published previously [6]. Apart from the predesigned mutation bands in rpoB coded gene, other mutations like L530P (n=2, 9.1%), S531F (n=6, 27.2%), S531W (n=10, 45.5%), A532V (n=2, 9.1%), and L533P (n=2, 9.1%) were observed in sequencing and these were also found resistant to RIF in pDST. In previous studies, disputed mutation at position L533P has been reported on both Lowenstein-Jensen and MGIT-960 culture medium [18,19]. Two specimens were sensitive to both sequencing and pDST. No MUT band was observed in LPA of these samples possibly due to failure of hybridization probe. The similar explanation has been quoted by Seifert et al. (2016) [10]. These patients’ isolates could have missed the proper resistant patterns to start the required treatment with single diagnostic test alone. This type of mutation frequencies was reported in ≥10% of total mutations associated with rpoB gene [19]. Mutation at codon 533 (L533P) is likely to be missed by Xpert MTB/RIF [20], but usually detected in LPA V.2, as seen in present study also (L533P [n=2, 9.1%]). LPA V.2 covers more coverage regions of rpoB gene (from codon 505 to 534) when compared to Xpert MTB/RIF (codon 509 to 533) [20]. However, some other deletions were also noted within and out of the hot spot region of rpoB gene which were associated with low level resistant in pDST [21,22]. Sequencing aided the identification of some other polymorphism in rpoB gene also. This finding of our study underscores that uncommon and/or silent mutation can cause NI results which can cause therapeutic dilemmas. Similarly, Nikam et al. [23] observed NI results in both RIF and INH without mentioning any specific loci of genes in MTBDRplus assay. Their study also highlighted this limitation of LPA which is especially of concern in TB endemic areas.
Therefore, this study highlights the limitations of LPA in detection of mutation patterns associated WT8 region (codon 530–533) [7]. To overcome this limitation, pDST and sequencing are recommended in select cases, such as NI results from LPAs. In the present study, we showed the usefulness of pDST and sequencing for elucidating the NI results from the Genotype MTBDRplus assay.
In most real-life scenario in high TB burden areas, sequencing may not be feasible. pDST can be a reasonable alternative in resource-limited settings. However, more prospective studies with clinical correlation can throw more light in this regard.
In the present study, we found that a small proportion of clinical samples were reported as NI in the Genotype MTBDRplus assay. Almost all of the strains with NI results (22/24, 91.7%) were RR in the pDST. Sequencing detected various rpoB mutations in these 22 cases. These results suggest that pDST and rpoB sequencing are useful in elucidating the cause and determining the clinical meaning of NI results from the Genotype MTBDRplus assay.
Notes
Authors’ Contributions
Conceptualization: Singh BK, Sharma R, Soneja M. Methodology: Singh BK, Sharma R, Soneja M, Jorwal P. Formal analysis: Singh BK, Sharma R. Software: Singh BK, Sharma R. Validation: Singh BK, Sharma R, Soneja M, Ramachandran R, Nischal N, Biswas A, Sarin S, Wig N. Investigation: Singh BK, Sharma R, Soneja M, Jorwal P, Nischal N. Writing – original draft preparation: Singh BK, Sharma R. Writing - review and editing: Soneja M, Kodan P, Nischal N, Biswas A, Sarin S, Wig N. Approval of final manuscript: all authors.
Conflicts of Interest
No potential conflict of interest relevant to this article was reported.
Funding
No funding to declare.
Acknowledgements
We are very thankful to Central TB Division, Ministry of Health & Family Welfare, Govt. of India; State TB Cell, National Capital Territory (NCT) of Delhi, Delhi; Foundation For Innovative New Diagnostics (FIND) India for logistic and technical support. Authors acknowledge support of lab personnel of Department of Medicine in carrying out this study